4-Aminobenzaldehyde: A Novel Antibiotic Based on 4-Aminobenzoic Acid and trans-Cinnamaldehyde
- Pre-Collegiate Global Health Review

- Apr 23, 2021
- 8 min read
By Om Pargaonkar, BASIS Scottsdale, Scottsdale, Arizona, USA
ABSTRACT
Antibiotic resistance is undoubtedly one of the greatest threats to humanity, threatening to undo nearly a century of global health advances. Antibiotics were once touted as wonder drugs that could cure even the gravest of bacterial infections. Now, faced with multi-drug resistant superbugs, they have been reduced to mere temporary solutions. To address this global crisis, scientists have looked to nature for novel alternatives. Several natural substances have been proven as effective antibiotics, including garlic, tea tree, cinnamon, and thyme extracts. After performing a Kirby-Bauer disk diffusion test, it was determined that cinnamon oil is the most effective of the four aforementioned natural extracts against the Gram-negative model bacterium, E. coli. This project thus combines the functional group of trans-Cinnamaldehyde, the primary antibiotic ingredient of cinnamon oil, with a molecule specifically taken up by bacteria, 4-Aminobenzoic acid, to produce 4-Aminobenzaldehyde. This molecule targets bacterial cells while remaining nontoxic to mammalian cells. 4-Aminobenzaldehyde is theorized to work by binding to and disrupting N-Acetylmuramic acid, a major constituent of peptidoglycan and thus bacterial cell walls. Through the development of this novel, naturally-derived antibiotic, this project demonstrates that the scientific community should seriously consider investigating natural antibiotic substances, determining their properties, and applying their functionality to bacteria-specific molecules to produce novel, effective antibiotic compounds.
INTRODUCTION
Antibiotic resistance is one of the greatest threats to humanity, threatening 2.8 million lives annually in the United States alone. Hundreds of strains of bacteria have emerged that are resistant to at least one antibiotic, causing infections that are particularly difficult to treat (C Reygaert, 2018). A prominent example of this is Methicillin-resistant Staphylococcus aureus (MRSA). This strain, which first emerged in the early 1960s, is credited with causing more annual infections and deaths than other seemingly more commonplace diseases such as AIDS (Green et al., 2012). To combat this looming crisis, scientists have continued attempting to synthesize new drugs. In fact, as of January 2020, there were 60 new antibiotic agents in some phase of testing (Bagozzi et al., 2020). Although this number may seem promising at first, it becomes much less so given that the majority of these products are simply new combinations of existing drugs. This lack of innovation at times has driven scientists to explore natural substances for antibiotic properties. Several experts have already established natural extracts as a potent source of novel antibiotic compounds, including Rossiter et al. who demonstrated the potential of several classes of natural substances as platforms for antibiotic development (Rossiter et al., 2017). However, one issue that is frequently encountered is the toxicity of some natural substances to mammalian cells, rendering several natural extracts impractical for antibiotic usage (Silver, 2011). Hence, the goal of this research is to determine a highly-effective natural antibiotic, analyze the functionality of the components of the natural extract, and apply the information to the development of a novel, naturally-inspired antibiotic compound that is largely nontoxic to mammalian cells (or, conversely, specific to bacterial cells).
METHODS
First, various natural extracts were tested to determine a highly effective natural antibiotic. For three of the natural extracts to be tested (T. vulgaris/Red thyme/Th, C. verum/Cinnamon/C, and M. alternifolia/Tea tree/TT), pure essential oils of the same brand were obtained. For the A. sativum (garlic/G) extract, whole bulbs were mashed, left to sit for 10 minutes to allow for allicin formation, and squeezed to extract the liquid. The garlic extract was used within 2 hours for maximum efficacy.

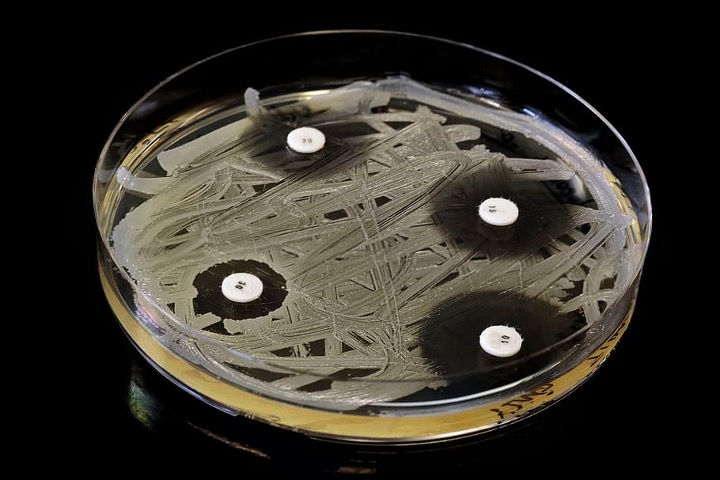
The extracts were tested using a Kirby-Bauer disk diffusion test. Agar plates were prepared for testing by pouring sterile Petri dishes with nutrient agar. After allowing the plates to properly set, they were inoculated with E. coli K12, following aseptic technique. 2.5-centimeter disks were cut out of Whatman #1 filter paper and soaked with equal volumes of test substances (1.5 μL). Lastly, the soaked filter paper disks were placed on the cultured agar plates and the plates were incubated at 37°C for 24 hours. Following incubation, the zone of inhibition (ZOI) diameter was measured for each test disk to determine efficacy. Once the most effective natural extract was identified, its active compound and activity pathway were identified through a review of relevant literature.
After completing initial testing to determine a natural compound to focus on, a modifiable molecule that is selectively taken up by bacteria was identified for use as a base for antibiotic development. Once the natural and base compounds were identified, a novel antibiotic was developed by combining the functional group of the natural compound with the essential group of the base compound. The structure was then refined by applying organic chemistry bonding principles to ensure maximum membrane permeability and efficacy against the target structure.
RESULTS
Most natural extracts were effective, with ZOI diameters of at least 10 ± 2 mm. C. verum extract proved to be the most effective with a ZOI diameter of 46.5 ± 1.5 mm.
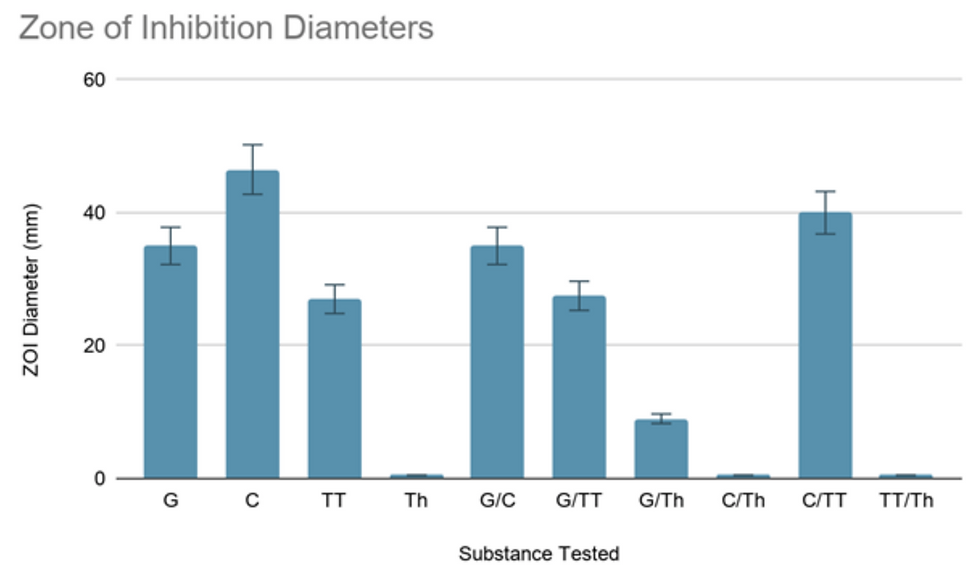
Fig. 2: A bar graph of the ZOI diameters.

Fig. 3: ZOI diameters of various natural extracts and combinations thereof.
Since cinnamon extract was proven to be the most effective, further research was conducted to identify active compounds and their action mechanisms. It was determined that trans-Cinnamaldehyde (tC) is the primary active ingredient of cinnamon essential oil, and that the molecule works by lowering intracellular ATP concentrations and disrupting the cell membrane and wall (Gill & Holley, 2004). The efficacy of tC is due to its aldehyde functional group, low relative polarity, and high membrane permeability (Gill & Holley, 2004). The functional group and traits of tC were recreated in the developed antibiotic.
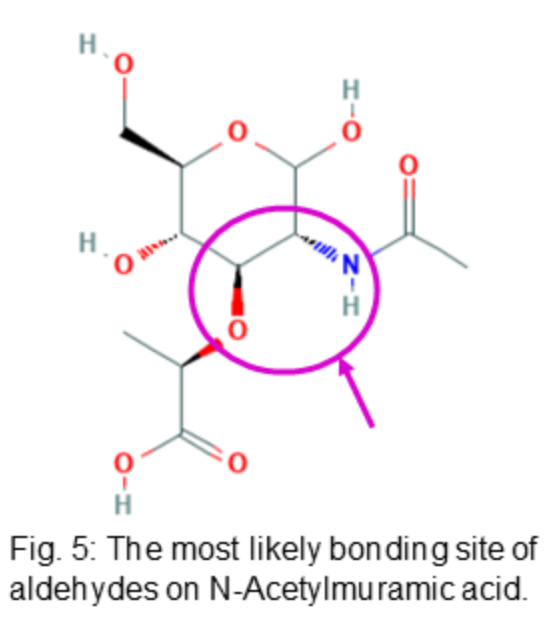
tC can target peptidoglycan, the main component of all bacterial cell walls. As such, it was chosen as the primary target for development of the antibiotic. The most probable bonding site of aldehyde-containing molecules was determined by applying organic chemistry principles. Specifically, it was found that aldehyde-containing molecules are most likely to bind to the nucleophilic peptide chain linkage site attached to N-Acetylmuramic acid (MurNAc), a major component of peptidoglycan. Thus, it is theorized that aldehyde-containing molecules would alter the attachment of the pentapeptide chain to MurNAc and disrupt the rigid structure of the bacterial cell wall.
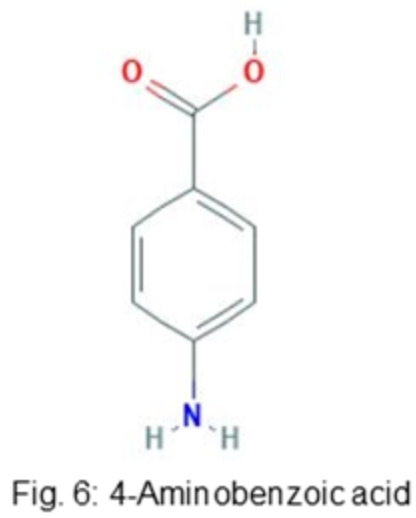
After researching a suitable base molecule for development of the antibiotic, 4-Aminobenzoic acid (pABA), a molecule selectively uptaken by bacteria for use in the folate synthesis pathway, was selected. The molecule pABA’s editable R-group (COOH) is attached to an aniline that is recognized by bacterial cells, making it a desirable candidate for antibiotic research (Mutch et al., 2018). Additionally, pABA is ignored by mammalian cells, which cannot utilize the substance (Mutch et al., 2018). The antibiotic sulfanilamide, which works as a competitive inhibitor of pABA in bacterial cells, is based on the structure of pABA, where the essential aniline group is maintained but the R-group is replaced with a sulfonyl amide group (Yun et al., 2012). Thus, this project altered pABA for use as an antibiotic, not only for its biochemical properties, but also for the similarity in structure between pABA and tC that will allow for straightforward combination of the functionality of the molecules.

Fig. 7: The antibiotic construction and refinement process.
The initial antibiotic molecule proposed after replacing the R-group of pABA with the functional group of tC was 4-Aminocinnamaldehyde. However, there are two key issues with this molecule: it is not as nonpolar/membrane-permeable as tC and it is less effective in binding to the R-O-R in MurNAc compared to tC due to its increased size and degrees of freedom. Thus, the molecular structure was refined by removing the C2H2 group between the aldehyde and 6-C ring on 4-Aminocinnamaldehyde. This change yielded the final proposed antibiotic molecule, 4-Aminobenzaldehyde. This molecule blends all of the properties desirable in an antibiotic: high selectivity for bacterial cells, low toxicity to mammalian cells, high membrane permeability, and high efficacy against the target structure, as determined by my research. 4-Aminobenzaldehyde should be perfectly equipped to travel in the bloodstream, selectively enter bacterial cells, and disrupt the cell wall by altering the attachment of the pentapeptide chain to MurNAc.
CONCLUSION
The development of a novel, broad-spectrum antibiotic is a major asset in the face of an antibiotic resistance crisis, especially given that billions of dollars are invested in antibiotic development every year. This project demonstrates, through the structural development of the antibiotic compound 4-Aminobenzaldehyde, that it is possible to alter known bacteria-specific molecules to possess antibiotic activity by utilizing the functionality of natural compounds. Additionally, this research highlights the potential of both pABA and aldehyde-containing molecules in future antibiotic research, which should also place a much greater emphasis on exploring natural substances for antibiotic properties.
My future research will focus on testing 4-Aminobenzaldehyde for in vivo toxicity against a model vertebrate, in vitro and in vivo antibiotic activity, and interactions with common drugs. Unfortunately, with the current circumstances related to the COVID-19 pandemic, I was not able to perform any of these tests due to the regulated laboratory setting they would have required. After proper experimentation and clinical trials, 4-Aminobenzaldehyde may be useful as a general antibiotic. Even if it is deemed unfit for internal infection treatment, it may be useful for the rapid sterilization of medical equipment in hospital environments.
Ultimately, the scientific community must rapidly turn its attention to the pressing matter of antibiotic resistance. It was recently demonstrated, in the development of COVID-19 vaccines, what the scientific community is capable of accomplishing in an extremely short period of time if its full potential is geared towards a common objective. This project was a humble contribution towards this ultimate goal. I am hopeful, for the sake of people everywhere, that the scientific community as a whole can develop a solution in time to address the global health crisis that is antibiotic resistance.
References
Bagozzi, D., Jasarevic, T., & Chaib, F. (2020, January 17). Lack of new antibiotics
threatens global efforts to contain drug-resistant infections. https://www.who.int/news/item/17-01-2020-lack-of-new-antibiotics-threatens-global-efforts-to-contain-drug-resistant-infections
C Reygaert, W. (2018). An overview of the antimicrobial resistance mechanisms of
bacteria. AIMS Microbiology, 4(3), 482–501. https://doi.org/10.3934/microbiol.2018.3.482
Gill, A. O., & Holley, R. A. (2004). Mechanisms of Bactericidal Action of
Cinnamaldehyde against Listeria monocytogenes and of Eugenol against L. monocytogenes and Lactobacillus sakei. Applied and Environmental Microbiology, 70(10), 5750–5755. https://doi.org/10.1128/aem.70.10.5750-5755.2004
Green, B. N., Johnson, C. D., Egan, J. T., Rosenthal, M., Griffith, E. A., & Evans, M. W.
(2012). Methicillin-resistant Staphylococcus aureus: an overview for manual
therapists. Journal of Chiropractic Medicine, 11(1), 64–76 https://doi.org/10.1016/j.jcm.2011.12.001
Green, D. W. (2002). The bacterial cell wall as a source of antibacterial targets. Expert
Opinion on Therapeutic Targets, 6(1), 1–20. https://doi.org/10.1517/14728222.6.1.1
Jean, N., Bougault, C., & Simorre, J.-P. (n.d.). GlycoPedia. Www.glycopedia.eu.
Retrieved February 27, 2021, from http://www.glycopedia.eu/e-chapters/the-structure-of-bacterial-cell/article/peptidoglycan-molecular-structure
Kapoor, G., Saigal, S., & Elongavan, A. (2017). Action and resistance mechanisms of
antibiotics: A guide for clinicians. Journal of Anaesthesiology Clinical Pharmacology, 33(3), 300. https://doi.org/10.4103/joacp.joacp_349_15
Kim, S., Chen, J., Cheng, T., Gindulyte, A., He, J., He, S., Li, Q., Shoemaker, B. A.,
Thiessen, P. A., Yu, B., Zaslavsky, L., Zhang, J., & Bolton, E. E. (2019).
PubChem in 2021: new data content and improved web interfaces. Nucleic acids research, 49(D1), D1388–D1395. https://doi.org/10.1093/nar/gkaa971
Mutch, C. A., Ordonez, A. A., Qin, H., Parker, M., Bambarger, L. E., Villanueva-Meyer,
J. E., Blecha, J., Carroll, V., Taglang, C., Flavell, R., Sriram, R., VanBrocklin, H.,
Rosenberg, O., Ohliger, M. A., Jain, S. K., Neumann, K. D., & Wilson, D. M. (2018). [11C]PABA: A PET tracer targeting bacteria-specific metabolism. ACS Infectious Diseases, 4(7), 1067–1072. https://doi.org/10.1021/acsinfecdis.8b00061
Rossiter, S. E., Fletcher, M. H., & Wuest, W. M. (2017). Natural Products as Platforms
To Overcome Antibiotic Resistance. Chemical Reviews, 117(19), 12415–12474. https://doi.org/10.1021/acs.chemrev.7b00283
Silver, L. L. (2011). Challenges of antibacterial discovery. Clinical Microbiology
Reviews, 24(1), 71–109. https://doi.org/10.1128/CMR.00030-10
Yun, M.-K., Wu, Y., Li, Z., Zhao, Y., Waddell, M. B., Ferreira, A. M., Lee, R. E.,
Bashford, D., & White, S. W. (2012). Catalysis and sulfa drug resistance in
dihydropteroate synthase. Science (New York, N.Y.), 335(6072), 1110–1114. https://doi.org/10.1126/science.1214641
Note: All molecular structures were obtained from the NIH National Library of Medicine’s PubChem database (Kim et al., 2019).




Comments